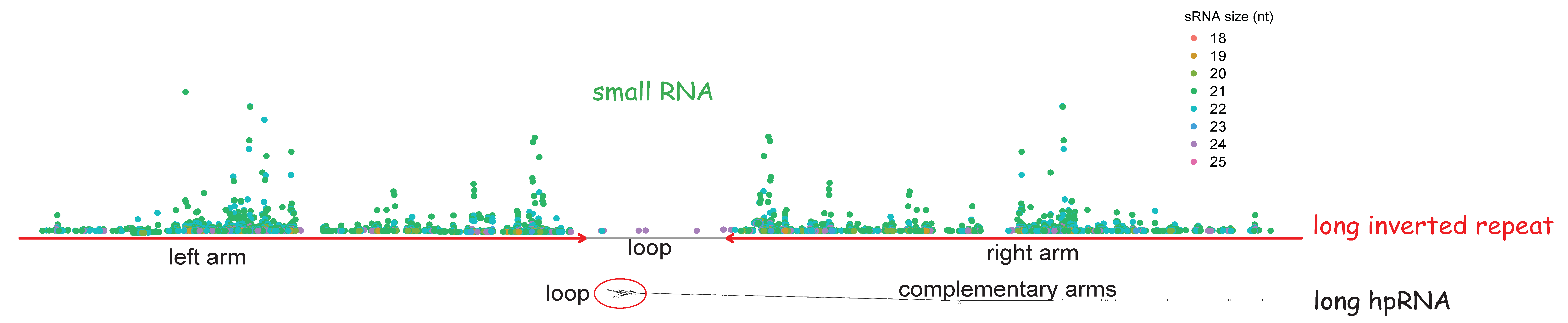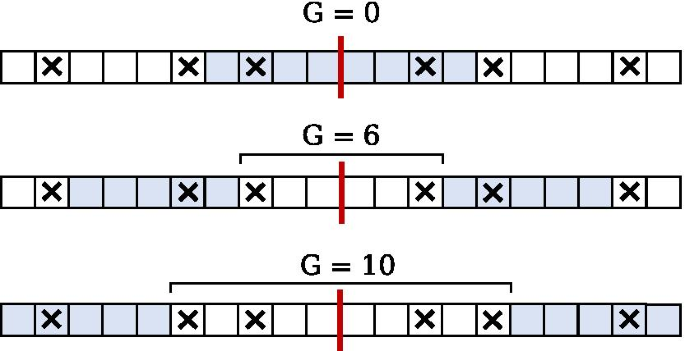
IUPACpal: efficient identification of inverted repeats in IUPAC-encoded DNA sequences | BMC Bioinformatics | Full Text
detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | PLOS ONE
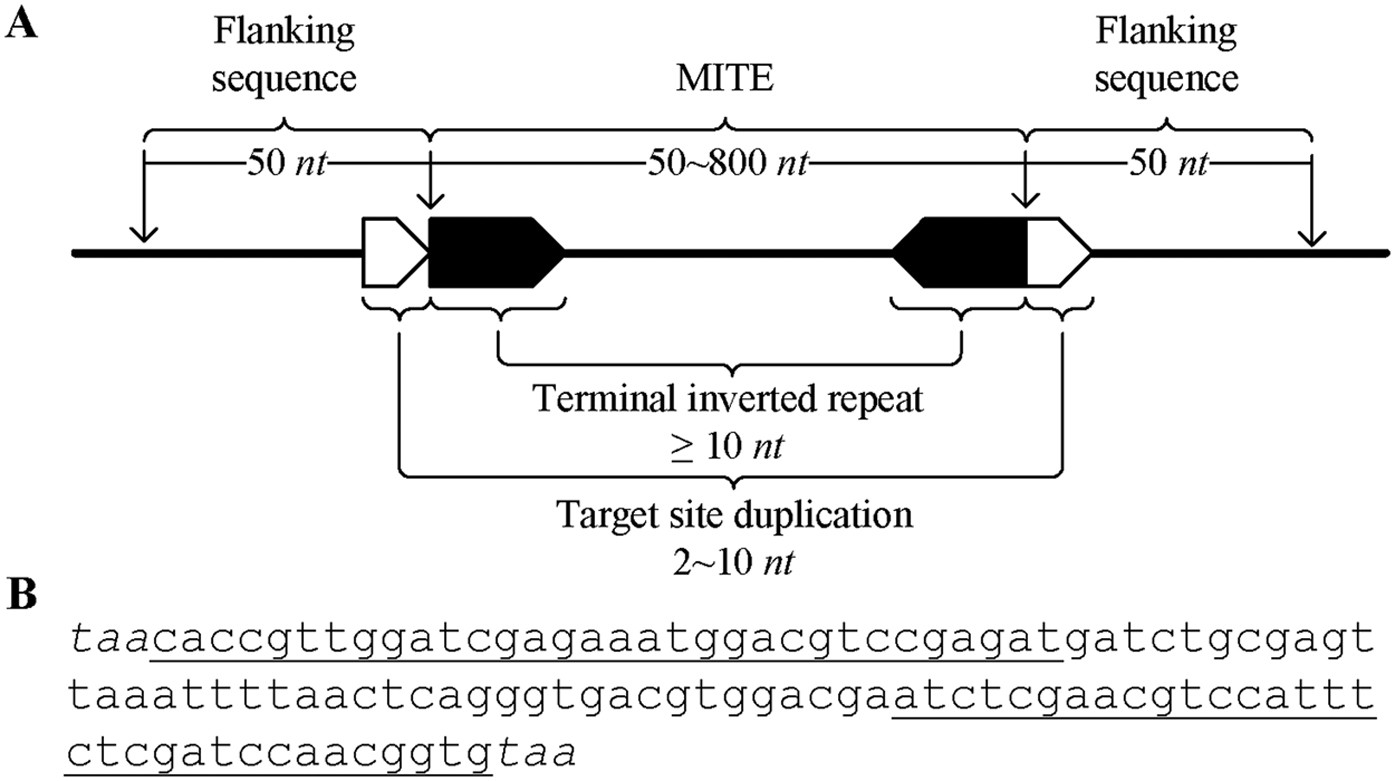
detectMITE: A novel approach to detect miniature inverted repeat transposable elements in genomes | Scientific Reports
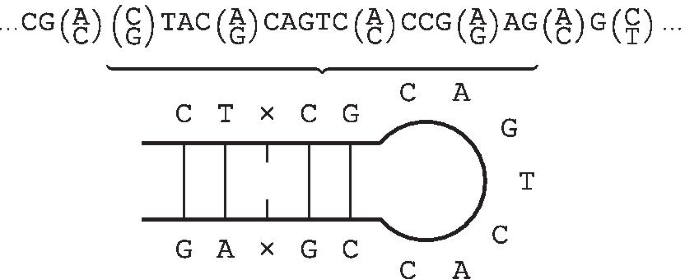
IUPACpal: efficient identification of inverted repeats in IUPAC-encoded DNA sequences | BMC Bioinformatics | Full Text

MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
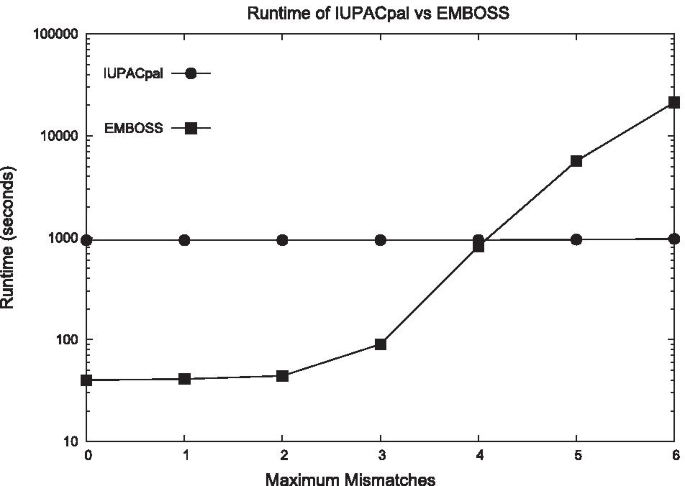
IUPACpal: efficient identification of inverted repeats in IUPAC-encoded DNA sequences | BMC Bioinformatics | Full Text

Generalized map of the chloroplast genome. IRA = inverted repeat region... | Download Scientific Diagram
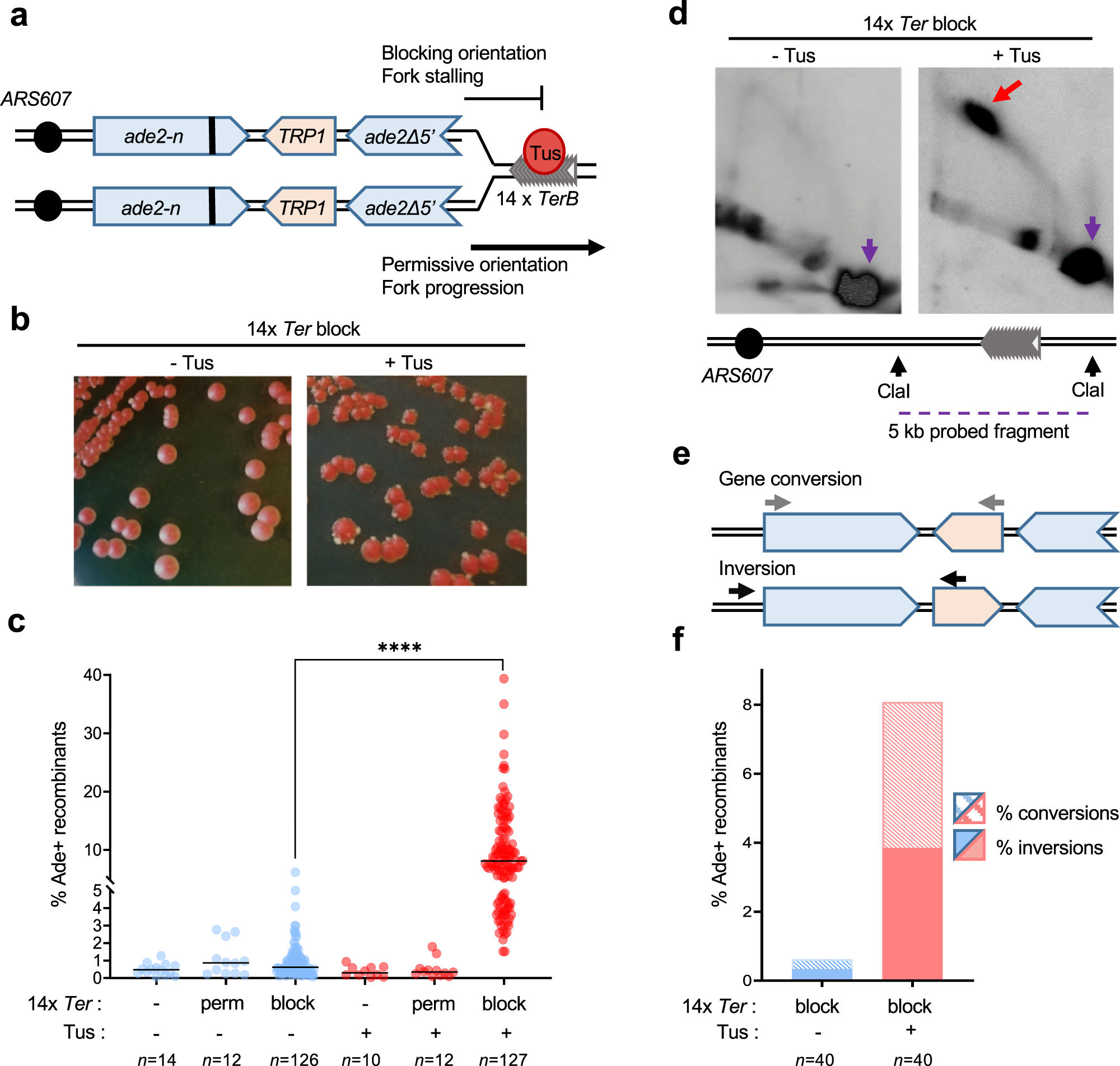
Mechanism for inverted-repeat recombination induced by a replication fork barrier | Nature Communications
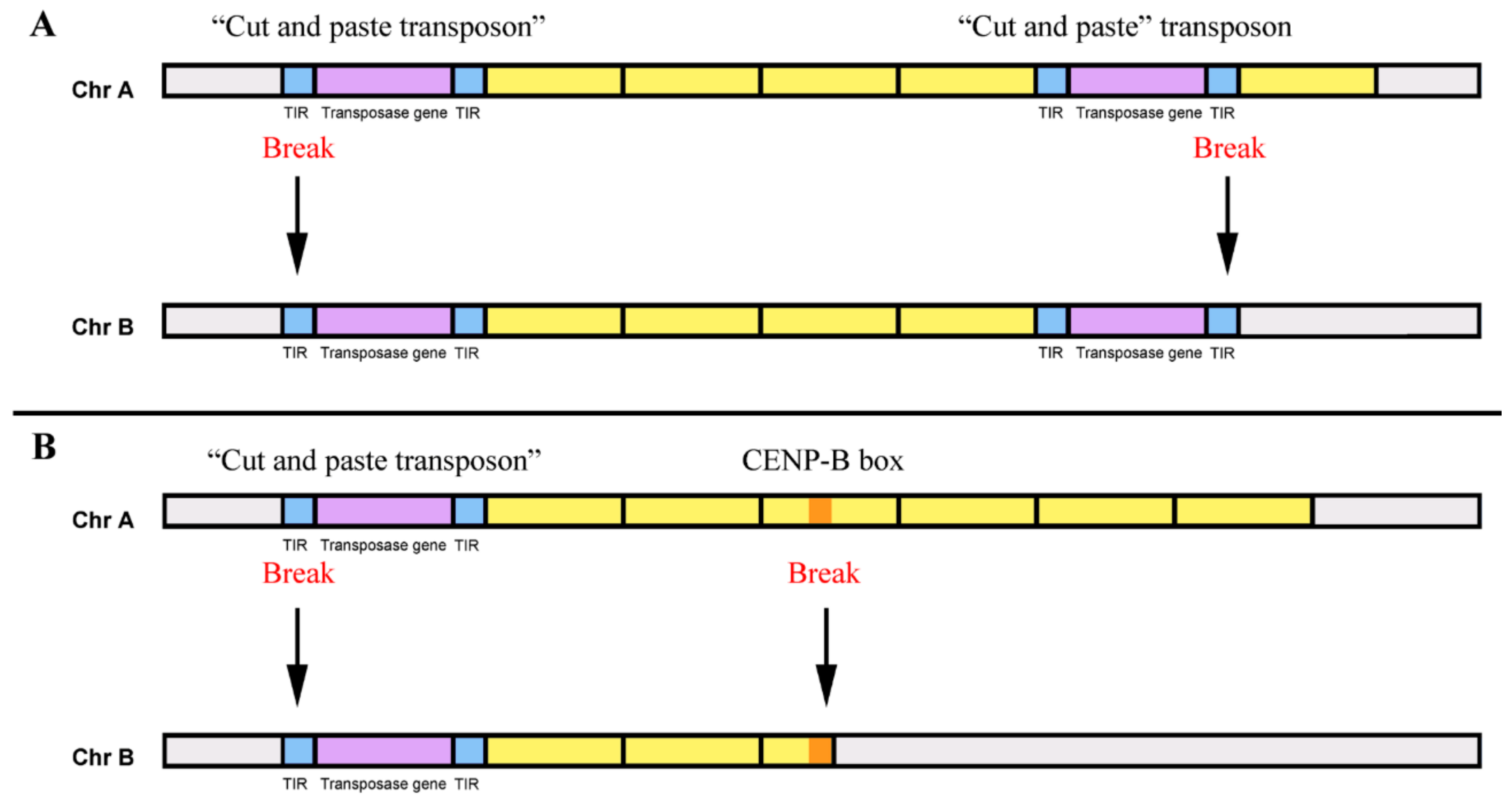
Genes | Free Full-Text | Conversion of DNA Sequences: From a Transposable Element to a Tandem Repeat or to a Gene